Abstract
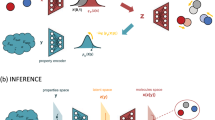
Optimizing molecular design and discovering novel chemical structures to meet specific objectives, such as quantitative estimates of the drug-likeness score (QEDs), is NP-hard due to the vast combinatorial design space of discrete molecular structures, which makes it near impossible to explore the entire search space comprehensively to exploit de novo structures with properties of interest. To address this challenge, reducing the intractable search space into a lower-dimensional latent volume helps examine molecular candidates more feasibly via inverse design. Autoencoders are suitable deep learning techniques, equipped with an encoder that reduces the discrete molecular structure into a latent space and a decoder that inverts the search space back to the molecular design. The continuous property of the latent space, which characterizes the discrete chemical structures, provides a flexible representation for inverse design to discover novel molecules. However, exploring this latent space requires particular insights to generate new structures. Therefore, we propose using a convex hull (CH) surrounding the top molecules regarding high QEDs to ensnare a tight subspace in the latent representation as an efficient way to reveal novel molecules with high QEDs. We demonstrate the effectiveness of our suggested method by using the QM9 as a training dataset along with the Self-Referencing Embedded Strings (SELFIES) representation to calibrate the autoencoder in order to carry out the inverse molecular design that leads to unfolding novel chemical structure.
This project is supported by the National Research Council Canada (NRC) and the Defence Research and Development Canada (DRDC).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only